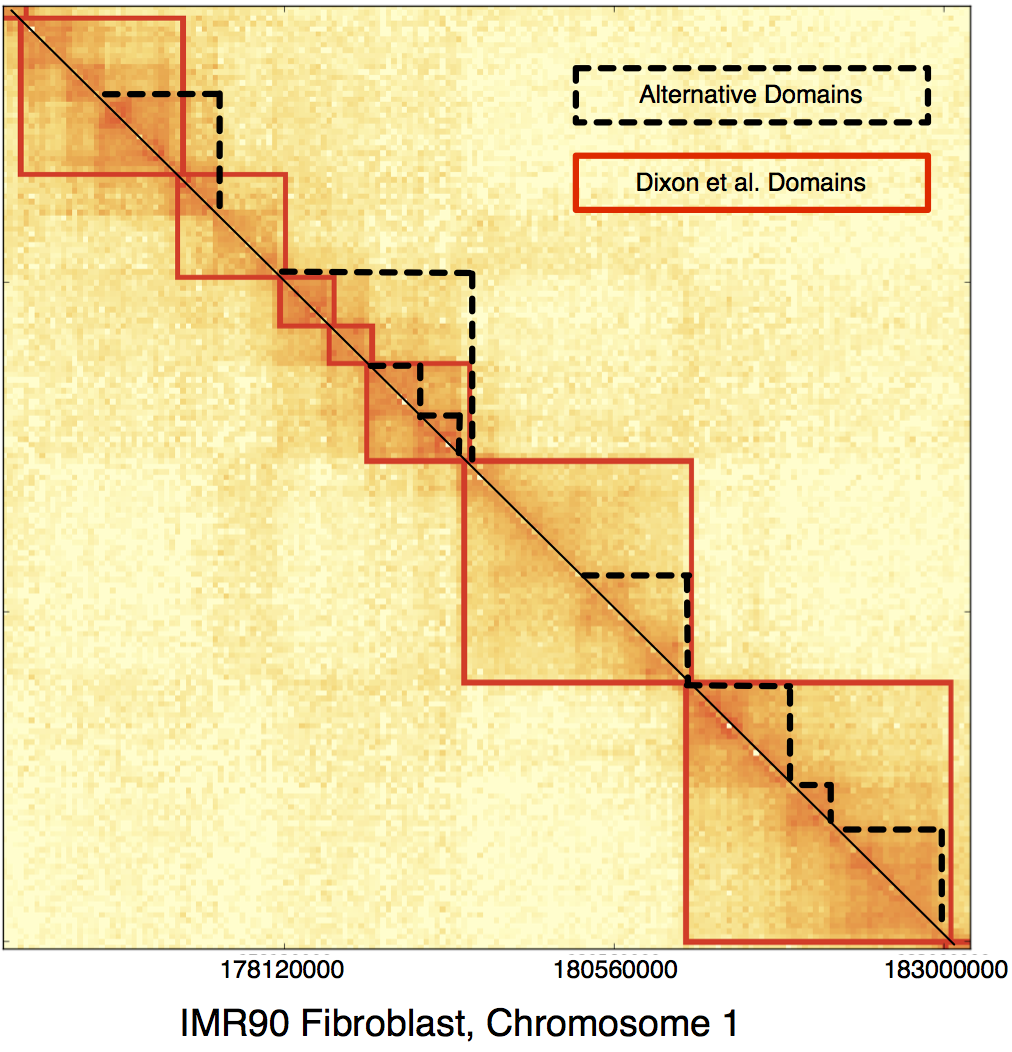
Recent chromosome conformation capture experiments have led to the discovery of dense, contiguous, megabase-sized topological domains that are similar across cell types and conserved across species. These domains are strongly correlated with a number of chromatin markers and have since been included in a number of analyses. However, functionally-relevant domains may exist at multiple length scales. We introduce a new and efficient algorithm that is able to capture persistent domains across various resolutions by adjusting a single scale parameter. The identified novel domains are substantially different from domains reported previously and are highly enriched for insulating factor CTCF binding and histone modfications at the boundaries.
[code]
[AMB version]
[Earlier WABI version]
Download domains:
- Armatus 2.0 consensus domain set (recommended)
Generated using C++ Armatus, commit ea54a41eade461283f154a19de8731d6de0be87c Changes since Armatus 1.0:- Changed denominator objective of equation 5 in the journal version from (l-k) to (l-k+1). In this definition, when l=k, the length of the domain is 1 as opposed to 0.
- For generating consensus domains with multiple optimal solutions, an ensemble of domains at a given resolution is weighted proportional to its objective function (normalized between 0 and 1).
- Armatus 1.0 consensus domain set (conference version + Java implementation):